Proteins tend to interact with surfaces and in doing so they can unfold and aggregate. The behavior of therapeutic proteins in solution is generally well characterized, but the adsorption to, and interaction with, various surface materials is often not studied. By studying protein interaction with various surface materials and at different solution conditions, the material compatibility can be evaluated and conditions that minimize adsorption can be identified. Here we show you one way to do this assessment.
Aggregation originating from protein-surface interaction could arise throughout all stages of drug manufacturing, storage, and distribution. Typical materials the protein would meet could be glass, metals, plastic polymers, and oils. The protein stability can however be enhanced by the addition of an excipient, such as a surfactant. So how does the addition of a surfactant affect the protein adsorption to a surface? Will the adsorption be prevented or just reduced? And would it be possible to identify a set of conditions that minimizes the amount of protein that adsorbs to the surface?
The nature of the protein adsorption depends on the specific combination of protein, surface, ambient conditions, and surfactant used. In the study here presented, the adsorption of two different monoclonal antibodies to four different surface materials was analyzed with QSense® QCM-D [1]. The adsorption was analyzed both with and without the presence of surfactant.
The questions to be answered were:

The QCM-D results, Fig. 1, show significant differences in the adsorbed amount, depending on which of the two antibodies that were measured, which surface material was used, and whether excipient was present or not.
Antibody: The amount of protein adsorbed varies with the antibody used. mAb2 shows a larger surface uptake than mAb1 on all four surface materials studied.
Surface material: The results show that the surface material significantly impacts the amount of protein adsorbed. The highest amount of adsorption is found on Au, and the lowest is found on PS.
Excipient: The presence of surfactant in the solution significantly reduces the amount adsorbed for both antibodies. For certain combinations of antibody and material, the adsorption is essentially prevented when the surfactant is added. The effect is most pronounced for mAb1 on PS.
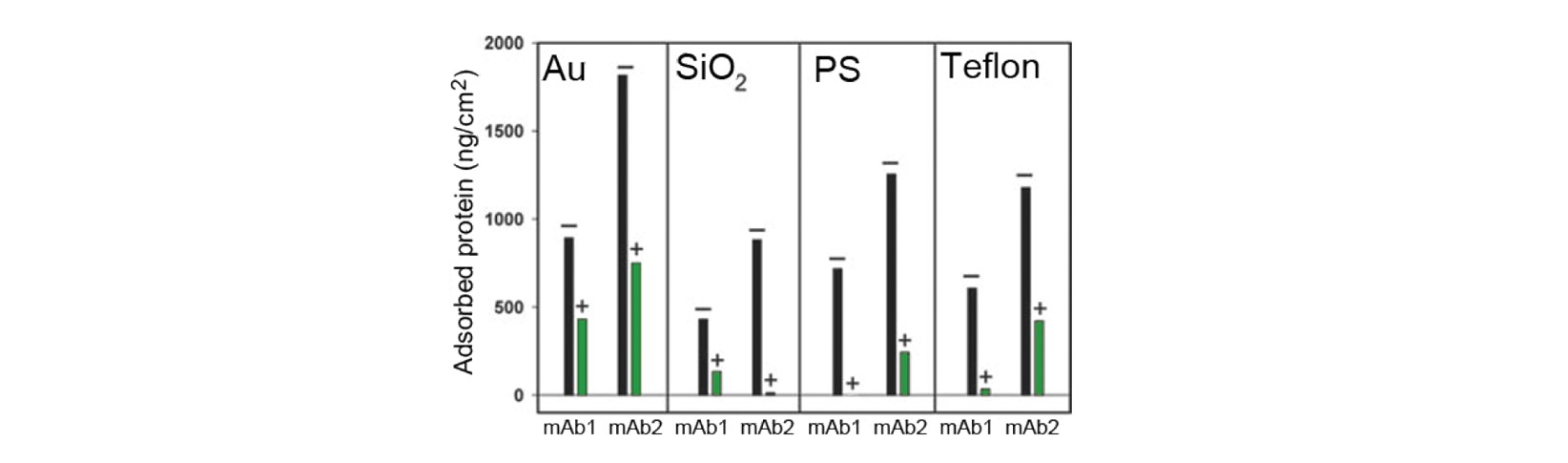
Figure 1. Mass of irreversibly adsorbed antibody to four different surface materials. + and – represent the presence (+) or absence (-) of surfactant [1].
Protein-surface interaction may trigger protein unfolding and aggregation and can result in loss of therapeutic properties as well as in immunogenic reactions. The extent of the protein-surface interaction depends on numerous factors, such as the nature of the protein itself, the surface material, pH, protein concentration, ionic strength, and presence of excipient, etc. In this study, the conditions that minimize protein-surface interaction within a defined context were identified via QCM-D analysis of mass uptake at the surface.
Download the case study to read more about the study.
Editor’s note: This post was originally published in May 2019 and has been updated.
Learn about how QSense analysis can be used to assess adsorption of biologics and excipients and reduce the risk of late discoveries of incompatibilities
Read about how protein adsorption at various surface and solution conditions quickly can be measured
Learn about how biointerfaces and biomolecular interactions can be studied using QSense® QCM-D and what information these measurements offer.
Read about how QSense QCM-D analysis is used in the quest to tackle inflammation and bacterial infections on implant surfaces.
Learn more about what questions QSense analysis can help answer in the context of container-closure interaction
Recent research results could help designing formulations better stabilized against protein aggregation, one of the main challenges in pharmaceutical development. QSense QCM-D analysis helped piecing the puzzle together
Read about how Dr. Gustaf Rydell, researcher at Sahlgrenska University Hospital, uses QCM-D to understand how the Norovirus initiates infection.
Here we present a list of scientific publications in virus-related research where QCM-D technology has been used
Learn more about how QSense QCM-D can be used to characterize biomolecular adsorption dynamics and the amount adsorbed to the surface of interest.
Here we present a summary of how protein adsorption to a silicone oil/water interface was analyzed in the presence and absence of surfactant.
Read about how molecule-surface interaction processes such as adsorption and desorption can be analyzed with QCM-D.
